Aluminum is the third most abundant element in earth’s crust, but its chemical properties have prevented its presence in the biological cycle of living organisms. Nevertheless, the acidification of the environment due mainly to human intervention has facilitated its solubilitation, thus increasing its bioavailability. Toxic effects of aluminum in the human body have been reported, and this element has been related with neurodegerentative diseases such as Alzheimer Disease. Aluminum has also been claimed to exert an important pro-oxidant activity.
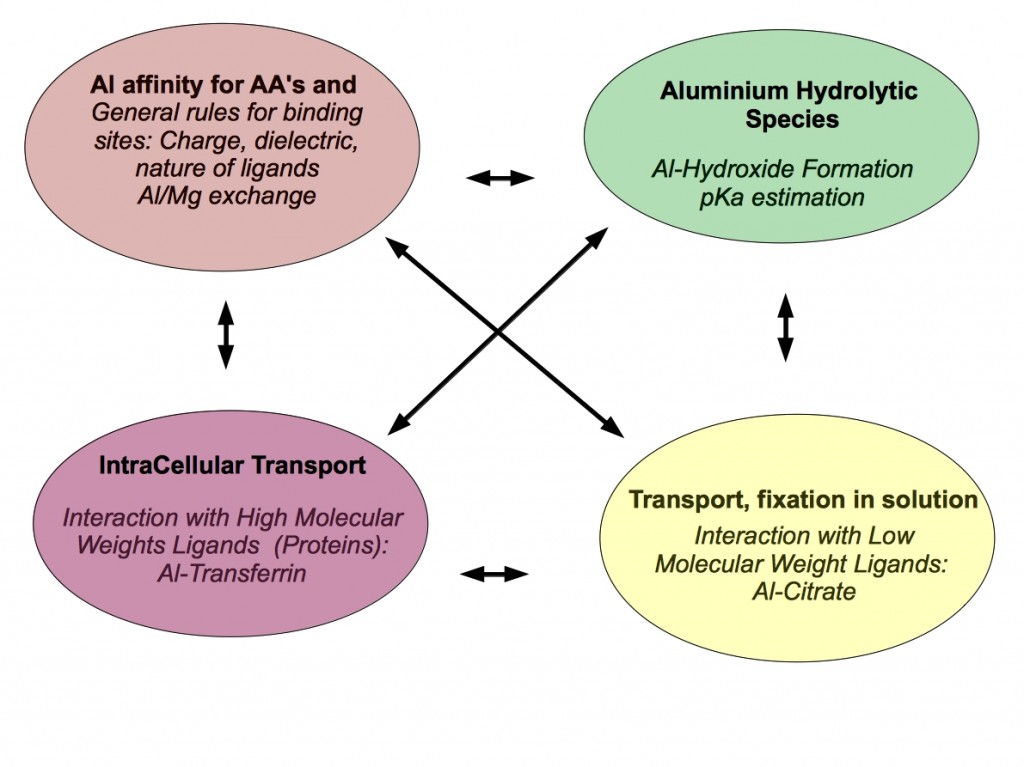
How aluminum can access the human organism is of primary importance. This implies knowledge about the kinetics, solubility and mobility of this element. In this sense, the speciation of aluminum has attracted the attention of many research groups. Aluminum is able to form a variety of hydrolysis products depending on the pH conditions, but Al(III) can also complexate with a variety of organic, inorganic or phosphate ligands. Of particular importance is the speciacion of aluminum in serum, where Al(III) interacts with high molecular mass (HMM) and low molecular mass (LMM) species. Among the HMM species the most important one is serum-Transferrin, and citrate and phoshates are among the main LMM chelators. In fact, it has been assumed, not without controversy, that aluminum can follow the iron pathway to enter the cell by receptor mediated endocytosis. Nevertheless, the interaction between Al(III) load transferrin and the receptor is questionable. Alternatively, it has been proposed that aluminum can get the brain not by Tf-TFR pathway, but rather complexated to citrate.
Whatever is the pathway used by Al(III) to enter in the cell, it is well know that its presence can alter the cell biochemistry in different ways. On one hand, it can disrupt metalloproteins that contain metal ions of similar size, such as Mg(II). Mg(II)/Al(III) substitution can thus alter the electronic and structural characteristics of the near-by residues, affecting the overall protein structure. On the other hand, Aluminun has demonstrated a significant pro-oxidant activity, which is still far from being understood at the molecular level.
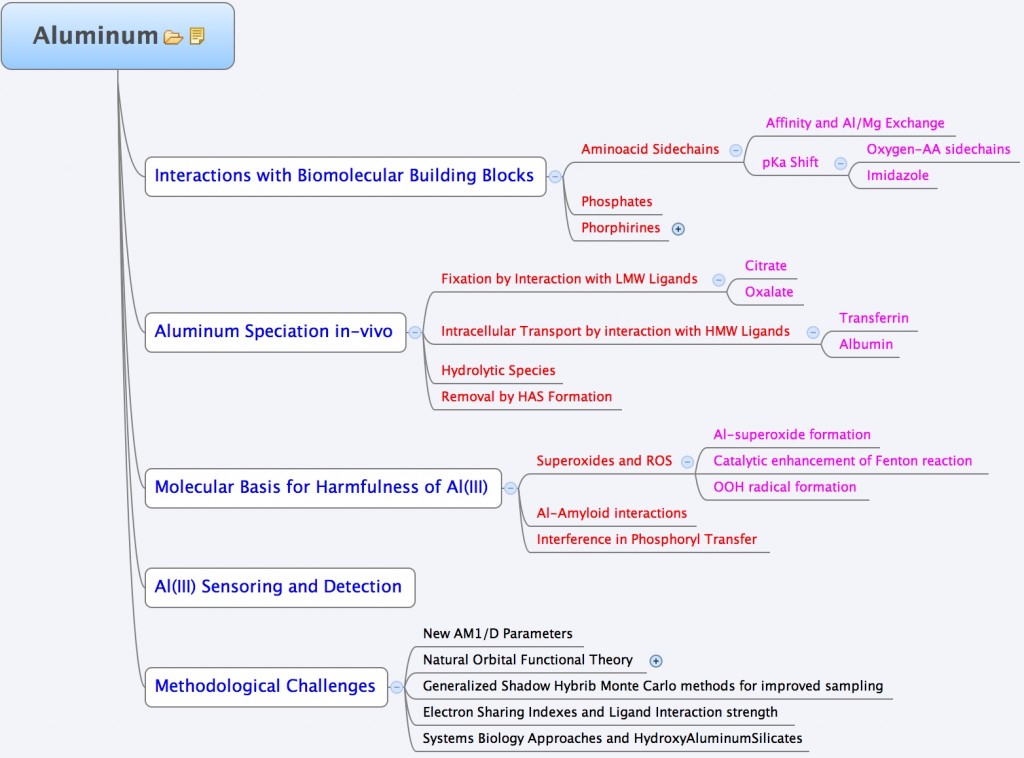
In the group, we use a variety of theoretical tools ranging from highly-accurate quantum chemical methods, to Force-Field Molecular dynamic simulations to unveil some of the aspects of the Aluminum interaction with biological systems. The following chart summarizes the different interests of the group in this area.
Publications of the group in this area:
- J. I. Mujika, J. Rodriguez-Guerra, X. Lopez, J. M. Ugalde, L. Rodriguez-Santiago, M. Sodupe, J. D. Maréchal, “Elucidating the 3D structures of Al(III)-Aβ complexes: a template free strategy based on the pre-organization hypothesis“, Chem. Sci., vol. 8, pp. 5041-5049 (2017), DOI: 10.1039/C7SC01296A
- E. Formoso, X. Lopez, “A computational study on interaction of aluminum with D-glucose 6-phosphate for various stoichiometries“, RSC Adv., vo. 7, pp. 6064-6079 (2017), DOI: 10.1039/C6RA27037A
- J. Beardmore, X. Lopez, J. I. Mujika, C. Exley, “What is the mechanism of formation of hydroxyaluminosilicates?“, Sci. Rep., vol. 6, pp. 30913 (2016), DOI: 10.1038/srep30913
- R. Grande-Aztatzi, E. Formoso, J. I. Mujika, J. M. Ugalde, X. Lopez, “Phosphorylation Promotes Al(III) Binding to Proteins: GEGEGSGG as a case study”, Phys. Chem. Chem. Phys., vol. 18, pp. 7197-7207 (2016), DOI:
- E. Formoso, J. I. Mujika, S. J. Grabowski, X. Lopez, “Aluminum and its effect in the equilibrium between folded/unfolded conformation of NADH“, J. Inorg. Biochem., vol. 152, pp. 139-146 (2015). DOI: 10.1016/j.jinorgbio.2015.08.017.
- N. B. Luque, J. I. Mujika, E. Formoso, X. Lopez, “Aluminum interaction with 2,3-diphosphoglyceric acid. A computational study“, RSC Adv. vol. 5, pp. 63874 (2015). DOI: 10.1039/c5ra06796k.
- N. B. Luque, Jon I. Mujika, E. Rezabal, J. M. Ugalde, X. Lopez, “Mapping the affinity of aluminum(III) for biophosphates: interaction mode and binding affinity in 1:1 complexes” Physical Chemistry Chemical Physics, v.16, pp.20107-20119 (2014). DOI: 10.1039/C4CP02770A
- J. I. Mujika, E. Rezabal, J. M. Mercero, F. Ruipérez, D. Costa, J. M. Ugalde, X. Lopez, “Aluminium in Biological Environments: A Computational Approach“, Computational and Structural Biotechnology Journal, vol.9, pp.1-13 (2014). DOI: 10.5936/csbj.201403002
- J. I. Mujika, J. M. Ugalde, X. Lopez, “Aluminum Interaction with Glutamate and α-Ketoglutarate. A Computational Study” The Journal of Physical Chemistry B, vol.118, iss.24, pp 6680–6686 (2014). DOI: 10.1021/jp502724w
- F. Ruipérez, J.I. Mujika, J.M. Ugalde, C. Exley, X. Lopez, “Pro-oxidant activity of Aluminum: Promoting the Fenton Reaction by reducing Fe(III) to Fe(II)“, Journal of Inorganic Biochemistry, (2012) .DOI: 10.1016/j.jinorgbio.2012.09.008
- J. I. Mujika, B. Escribano, E. Akhmatskaya, J. M. Ugalde, X. Lopez, “Molecular Dynamics Simulations of Iron- and Aluminum-Loaded Serum Transferrin: Protonation of Tyr188 Is Necessary To Prompt Metal Release“, Biochemistry, vol.51, iss.35, p.7017-7027, (2012) . DOI: 10.1021/bi300584p
- J. I. Mujika, J. M. Ugalde, X. Lopez, “Aluminum speciation in biological environments. The deprotonation of free and aluminum bound citrate in aqueous solution“, Physical Chemistry Chemical Physics, (2012) . DOI: 10.1039/C2CP40671C
- J.I. Mujika, X. Lopez, E. Rezabal, R. Castillo, S. Marti, V. Moliner, J.M. Ugalde, “A QM/MM study of the complexes formed by aluminum and iron with serum transferrin at neutral and acidic pH“, Journal of Inorganic Biochemistry, vol.105, iss.11, p.1446-1456, (2011) . DOI: 10.1016/j.jinorgbio.2011.07.019
- J. I. Mujika, F. Ruiperez, I. Infante, J. M. Ugalde, C. Exley, X. Lopez, “Pro-oxidant Activity of Aluminum: Stabilization of the Aluminum Superoxide Radical Ion“, The Journal of Physical Chemistry A, vol.115, iss.24, p.6717-6723, (2011) . DOI: 10.1021/jp203290b
- J. I. Mujika, J. M. Ugalde, X. Lopez, “Computational evaluation of pK a for oxygenated side chain containing amino acids interacting with Aluminum“, Theoretical Chemistry Accounts, vol.128, iss.4-6, p.477-484, (2010) . DOI: 10.1007/s00214-010-0807-6
- E. Rezabal, T. Marino, J. M. Mercero, N. Russo, J. M. Ugalde, “Complexation of Al“, Inorganic Chemistry, vol.46, iss.16, p.6413-6419, (2007) . DOI: 10.1021/ic7004776
- Elixabete Rezabal, Jose M. Mercero, Xabier Lopez, Jesus M. Ugalde, “A theoretical study of the principles regulating the specificity for Al(III) against Mg(II) in protein cavities“, Journal of Inorganic Biochemistry, vol.101, iss.9, p.1192-1200, (2007) . DOI: 10.1016/j.jinorgbio.2007.06.010
- Elixabete Rezabal, Jose M. Mercero, Xabier Lopez, Jesus M. Ugalde, “A study of the coordination shell of aluminum(III) and magnesium(II) in model protein environments: Thermodynamics of the complex formation and metal exchange reactions“, Journal of Inorganic Biochemistry, vol.100, iss.3, p.374-384, (2006) . DOI: 10.1016/j.jinorgbio.2005.12.007
- Elixabete Rezabal, Jose M. Mercero, Xabier Lopez, Jesus M. Ugalde, “Protein Side Chains Facilitate Mg/Al Exchange in Model Protein Binding Sites“, ChemPhysChem, vol.8, iss.14, p.2119-2124, (2007) . DOI: 10.1002/cphc.200700335
- Jose M. Mercero, Jon M. Matxain, Elixabete Rezabal, Xabier Lopez, Jesus M. Ugalde, “The first solvation shell of aluminum (III) and magnesium (II) cations in a protein model environment“, International Journal of Quantum Chemistry, vol.98, iss.4, p.409-424, (2004) . DOI: 10.1002/qua.20075
- Eusko Jaurlaritza (Basque Government): Ref: GIC 07/85 IT-330-07
- Ministerio de Ciencia e Innovacion (Spanish Government): CTQ 2011-27374
- EPSRC: EP/J004146/1
Collaborations with other groups:
- Christopher Exley, Keele University, United Kingdom
- Darrin M. York, Rutgers University, USA
- Eduard Matito, Universitat de Girona
- Elena Akhmatskaya, Basque Center for Applied Mathematics
- Mariona Sodupe, Universitat Autonoma de Barcelona
For more information contact xabier.lopez@ehu.es